MACE¶
MACE is a equivariant machine learning code for developing accurate machine learning interatomic potentials. The main references from the MACE architeture are 1 and 2. A lot of important information is also available on the code's repository and documentation.
Training¶
Finetuning¶
One of the main advantages of MACE is the ability to finetune foundation models on both your target dataset and a replay dataset derived from the foundation model. This approach helps prevent adapting the model to your specific dataset and reference level, while retaining the stability from the foundation model.
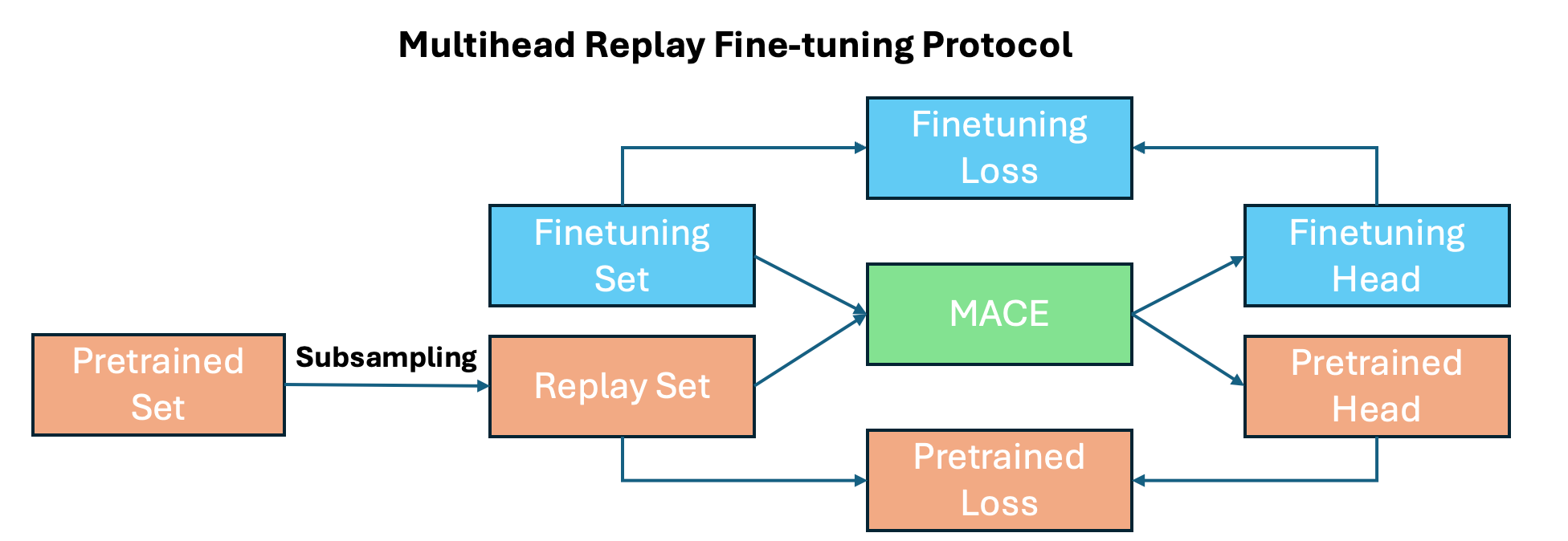
Get foundation model dataset¶
Obtain a replay dataset from the foundation model with
mace_finetuning \
--configs_pt <path-to-replay-data> \
--configs_ft <path-to-target-data> \
--num_samples <number-of-points> \
--subselect <fps/random> \
--model <path-to-foundation-model> \
--output <path-to-output-file> \
--filtering_type <combinations/exclusive/inclusive> \
--head_pt pt_head \
--head_ft target_head \
--weight_pt <1.0> \
--weight_ft <10.0>
Calculate isolated atoms (E0s)¶
Finetuning a ScaleShiftMACE model requires the isolated atom energies (E0s), that must be computed consistently:
- Use the same
POTCARandENCUTas the reference/training data. - Place the atom in a large cell (e.g. 20–30 Å box) slightly non-cubic cells sometimes converge more stably.
- Ensure correct electron occupation in OUTCAR, see example
- Add energies to training files with
config_type=IsolatedAtomor setE0s: <name-json-file>to the configuration file.
An example for how to prepare all this file "automatically" can be found here.
Quick finetune example¶
For finetuning you can just run mace_run_train --config ft.yml with a configuration file similar to this:
name: <name>
foundation_model: <path-to-foundation-model>
multiheads_finetuning: True
foundation_filter_elements: True
foundation_model_readout: True
loss: universal
heads:
ft_head:
train_file: <path-train-data>
valid_file: <path-validation-data>
test_file: <path-test-data>
E0s: IsolatedAtom
energy_key: dft_energy
forces_key: dft_forces
stress_key: dft_stress
pt_train_file: <path-to-replay-data>
energy_key: energy
forces_key: forces
stress_key: stress
atomic_numbers: "[1, 6, 8, 13, 14]"
foundation_model_elements: True
energy_weight: 1.0
forces_weight: 10.0
stress_weight: 10.0
model: "ScaleShiftMACE"
compute_stress: True
compute_forces: True
clip_grad: 10
amsgrad: True
error_table: PerAtomRMSE
weight_decay: 1e-8
eval_interval: 1
lr: 0.001
scaling: rms_forces_scaling
batch_size: 6
max_num_epochs: 512
ema: True
ema_decay: 0.9999
save_all_checkpoints: True
patience: 50
default_dtype: float64
device: cuda
restart_latest: True
seed: 2025
keep_isolated_atoms: True
save_cpu: True
enable_cueq: False
wandb: True
wandb_project: FeZeos
wandb_entity: nano-materials-modelling
wandb_name: FT_fezeo_matpes_500
Not all this keywords are required, you can find more information on the MACE documentation.
Foundation models¶
MACE MP0¶
MACE-MP0 models are foundation models trained using the Materials Project database. The models are available here. The second-generation models, MACE-MP-0b and MACE-MP-0b2, include the following improvements for simulations: - Enhanced stability via core repulsion. - Regularization for high-pressure conditions. - Additional training data at high-pressure.
For most use cases, we recommend using the second-generation models for MD simulations and fine-tuning, unless earlier models are explicitly required. They can be found on Metacentrum at /storage/praha2-natur/home/carlosbornes/MACE/models
MATPES¶
MD simulations¶
Using Janus¶
Janus is a Python package that interfaces with multiple machine learning potentials, such as MACE, M3GNet, CHGNet, and others. It can be used both in Jupyter notebooks and the terminal. Example notebooks and configuration files are available here.
You can find an example of a NVT MD simulation of Cu supported on alumina at /storage/praha2-natur/home/carlosbornes/MACE/Examples/MD
Quick explanation nvt.yml Configuration File¶
struct: POSCAR
ensemble: nvt
steps: 1000000
timestep: 1
temp: 600
arch: mace_mp
device: cuda
stats_every: 1000
traj-every: 1000
tracker: False
remove_rot: True
rotate_restart: True
calc-kwargs:
model: medium-0b2
enable_cueq: True
dispersion: True
skin: 2.0
check: True
delay: 1
every: 1
Key Parameters:¶
struct: The POSCAR file used for the simulation. Supports all formats compatible with ASE.ensemble: Specifies the ensemble type (e.g.,nvtfor constant volume and temperature).remove_rot: Removes rotational motion during the MD simulation. Particularly useful fornptsimulations.rotate_restart: Controls the frequency of saving restart files. Defaults toFalse; if set, saves every 1000 steps.calc-kwargs: Additional arguments passed to the calculator, including:model: Specifies the model used for the simulation (e.g.,medium-0b2).enable_cueq: Enables NVIDIA's cuEquivariance library for faster GPU performance.dispersion: Activates dispersion corrections for the model.skin: Sets the neighbor list buffer (skin) size to avoid recomputation at every step. Useful when dispersion corrections are enabled.
Additional Notes:¶
- To load Janus, use the following command:
- For a full list of MD-related options, run: ```bash janus md --help